What Are the Experimental Workflows in Golgi Apparatus Proteomics?
-
Differential centrifugation combined with density gradient centrifugation (e.g., sucrose gradients) which represent classical methods for effective organelle separation.
-
Commercial enrichment kits, such as the Golgi Enrichment Kit from Thermo Fisher Scientific, can enhance operational efficiency and reproducibility.
-
Immunomagnetic isolation (Immunoisolation), which employs antibodies targeting Golgi marker proteins (e.g., GM130) together with magnetic beads to achieve selective enrichment. This strategy improves purity but increases cost.
-
Strict temperature control should be maintained to minimize organelle degradation.
-
Isolation purity can be evaluated by Western blot analysis of markers for the Golgi (GM130) and the ER (Calnexin) to assess contamination levels.
The Golgi apparatus is a critical membranous organelle responsible for protein modification, sorting, and trafficking. Comprehensive characterization of the Golgi proteome facilitates understanding of its functional roles in the secretory pathway, glycosylation, and disease development. With advances in high-resolution mass spectrometry, Golgi proteomics has become a central strategy for investigating subcellular organelle function.
Sample Preparation and Golgi Isolation
1. Selection of Sample Types
Golgi proteomics investigations can be conducted using cell lines (e.g., HeLa, HepG2) or animal tissues (e.g., liver, brain). Cell lines are frequently selected because they provide consistent biological sources and are convenient to manipulate.
2. Golgi Enrichment and Purification
Because the Golgi apparatus is physically and functionally associated with membranous compartments such as the endoplasmic reticulum and lysosomes, achieving high purity is critical for experimental reliability. Commonly applied approaches include:
Notes:
Protein Extraction and Quantification
As the Golgi is a membrane-rich structure, many proteins are integral or membrane-associated. Therefore, suitable lysis conditions are required, including:
1. Buffers containing detergents such as Triton X-100 or NP-40.
2. High-salt or alkaline extraction protocols for peripheral membrane proteins.
Protein abundance can be determined using the BCA assay.
Proteolytic Digestion and Peptide Purification
1. Protease Digestion
Trypsin is routinely employed, and Lys-C may be introduced when necessary to improve proteolytic efficiency.
2. Peptide Purification and Desalting
C18 solid-phase extraction (SPE) columns are applied to remove salts and other contaminants, thereby preventing ionization suppression.
For modification-focused analyses (e.g., glycosylation), additional deglycosylation or enrichment procedures are required.
Mass Spectrometry Analysis (LC-MS/MS)
High-resolution mass spectrometry platforms, including Orbitrap Exploris and timsTOF Pro, enable accurate peptide characterization.
Widely used acquisition strategies include:
1. DDA (Data Dependent Acquisition), typically applied in discovery-driven studies.
2. DIA (Data Independent Acquisition), which enhances proteome coverage and quantitative robustness.
3. Isobaric labeling strategies such as TMT or iTRAQ, suitable for multi-group comparisons.
Integration with nano-flow liquid chromatography (nanoLC) is recommended to improve analytical sensitivity.
Data Analysis and Functional Annotation
1. Raw spectra are commonly analyzed using platforms such as MaxQuant and Proteome Discoverer for peptide identification, protein inference, and quantification.
2. Perseus is widely applied for downstream statistical evaluation, including differential analysis and clustering.
3. Functional interpretation and subcellular localization are supported by resources such as GO, KEGG, and SubCell Barcode.
Comparison with established Golgi marker proteins enables assessment of isolation success and verification of Golgi localization.
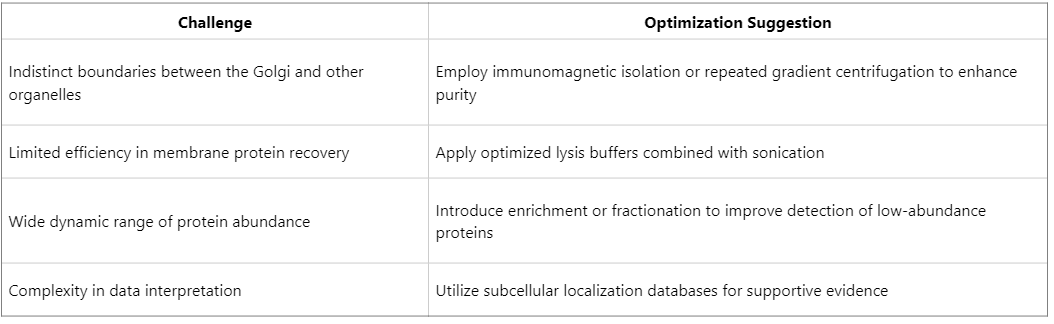
Challenges and Optimization Strategies in Golgi Proteomics
Service Advantages of MtoZ Biolabs in Subcellular Proteomics
At MtoZ Biolabs, advanced ultracentrifugation platforms, high-sensitivity mass spectrometry systems (e.g., Orbitrap Exploris 480), and proprietary subcellular localization data-analysis pipelines are integrated to support quantitative proteomics of organelles such as the Golgi, mitochondria, and lysosomes.
Key service features include:
1. Assessment of Golgi enrichment purity combined with multi-omics analysis.
2. Flexible implementation of labeled or label-free quantification strategies (DIA/TMT/SILAC).
3. Tailored experimental designs adaptable to disease models, pharmacological interventions, and related applications.
4. Comprehensive deliverables encompassing data analysis, functional annotation, and visualization.
Researchers may contact us to obtain customized solutions for Golgi proteomics.
Golgi proteomics provides a powerful framework for investigating dynamic alterations within the intracellular membrane system and has broad applications in cell biology, oncology, and neuroscience. Through rigorous workflow design and high-quality data generation, functional changes of the Golgi and their mechanistic relevance to disease can be elucidated in depth. For studies targeting subcellular proteomics, MtoZ Biolabs offers integrated technical support within a one-stop service model.
MtoZ Biolabs, an integrated chromatography and mass spectrometry (MS) services provider.
Related Services
How to order?







