Resources
Proteomics Databases
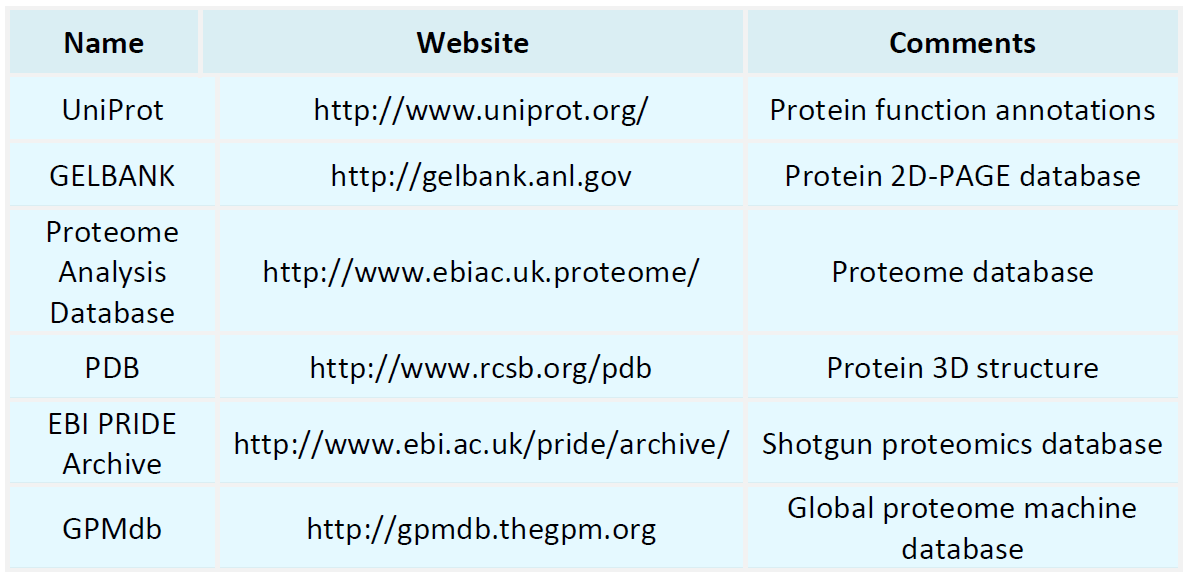
Metabolomics Databases
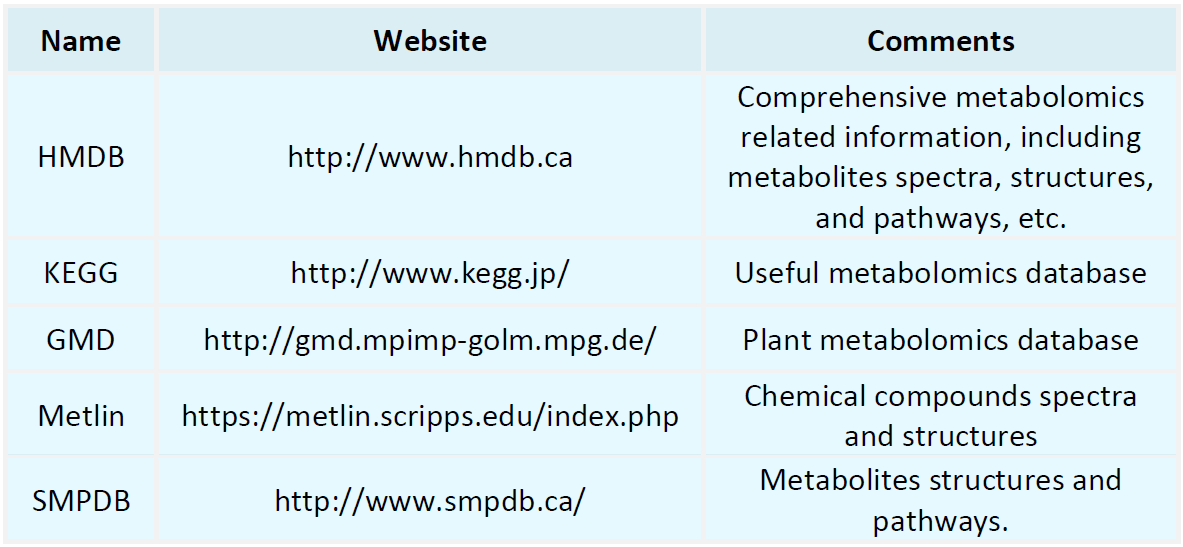
-
• Detection of Proteomic Changes Using SILAC/Dimethyl Labeling
In contemporary biological research, proteomics stands at the forefront of elucidating the functions and mechanisms of biological systems. Protein expression levels, post-translational modifications, and interactions are pivotal in cellular biology studies. Within proteomics, quantitative analysis serves as a vital approach to uncovering these molecular shifts.
-
• DIA-Based Detection and Analysis of Protein Biomarkers
In modern biomedical research, the detection and analysis of protein biomarkers are critical for diagnosis, treatment, and disease monitoring. Data-Independent Acquisition (DIA), an advanced mass spectrometry technique, is widely used in proteomics due to its high throughput, reproducibility, and broad data coverage.
-
• Phosphoproteomics Analysis Based on TMT Labeling
Phosphorylation is a crucial post-translational modification (PTM) that plays an essential role in controlling protein function, cellular signaling, and metabolic processes. Aberrant phosphorylation is often closely linked to the development and progression of various diseases. Therefore, comprehensive phosphoproteomic analysis is vital for understanding biological processes and disease mechanisms.
-
• Quantitative Analysis of Protein Binding Affinity Using Far-Western Blot
Protein-protein interactions (PPIs) are essential to biological processes such as cellular signaling, metabolic regulation, and immune responses. Quantitative analysis of protein binding affinity is critical for understanding these interactions' biological significance. Far-Western blotting, a specialized protein analysis technique, employs labeled probe proteins to detect interactions with target proteins, enabling the precise quantification of binding affinity.
-
• Analysis of Protein Complexes Using AP-MS
Protein complexes play essential roles in various cellular functions, including signal transduction, metabolic regulation, and gene expression. Understanding the composition and interactions of these complexes is crucial for unraveling cellular functions and elucidating disease mechanisms. In recent years, Affinity Purification-Mass Spectrometry (AP-MS) has emerged as a powerful technique widely used in protein complex research.
-
• Analysis of Protein Complexes Using Co-Immunoprecipitation and Mass Spectrometry
Proteins typically function within cells as part of complexes, and the assembly and dynamic changes of these complexes are crucial for understanding cellular signaling, metabolic pathways, and other biological processes. The combination of immunoprecipitation (Co-IP) with mass spectrometry (MS) is a powerful technique for studying and dissecting the composition and interaction networks of protein complexes.
-
• Application of PCT-DIA Proteomics
Proteomics is an essential tool in modern life sciences for studying protein expression, function, and interactions. Among the various proteomics techniques, Data-Independent Acquisition (DIA) based methods have become increasingly popular due to their high throughput, sensitivity, and quantitative accuracy. However, as proteomics research advances, there is a growing demand for efficiency and consistency in protein sample preparation.
-
• Mechanism of PCT-DIA Proteomics
Proteomics, the field dedicated to the study of the structure, function, and interactions of proteins within a biological system, is fundamental to advancing our understanding of life sciences. The integration of Pressure Cycling Technology (PCT) with Data-Independent Acquisition (DIA) represents a significant advancement in proteomics, enhancing both the efficiency and precision of analysis.
-
• Workflow of PCT-DIA Proteomics
PCT-DIA (Pressure Cycling Technology-DIA) is an advanced analytical approach in modern proteomics. By combining Pressure Cycling Technology (PCT) with Data-Independent Acquisition (DIA), this method facilitates efficient, comprehensive, and highly sensitive analysis of complex protein samples. PCT-DIA not only improves protein identification rates but also significantly enhances the accuracy of quantitative data, establishing itself as a pivotal tool in proteomics research.
-
• Advantages and Disadvantages of PCT-DIA Proteomics
PCT-DIA (Pressure Cycling Technology-DIA) is a proteomics analysis method that combines Pressure Cycling Technology (PCT) with Data-Independent Acquisition (DIA). This technique enhances protein lysis efficiency by alternating between high and low pressures, allowing for comprehensive protein quantification using DIA.
How to order?







