Resources
Proteomics Databases
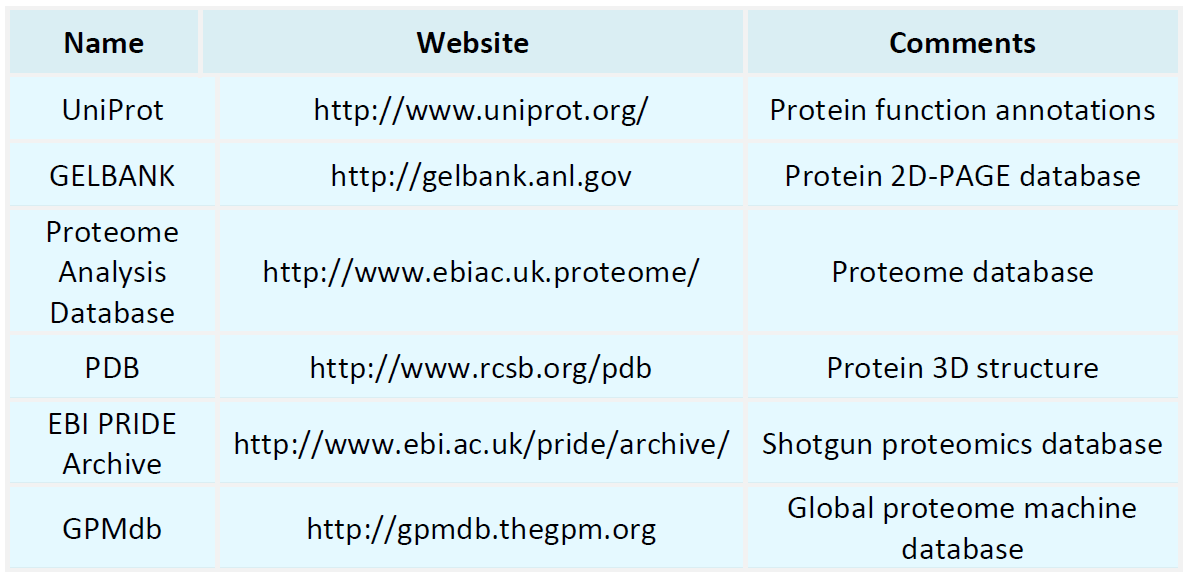
Metabolomics Databases
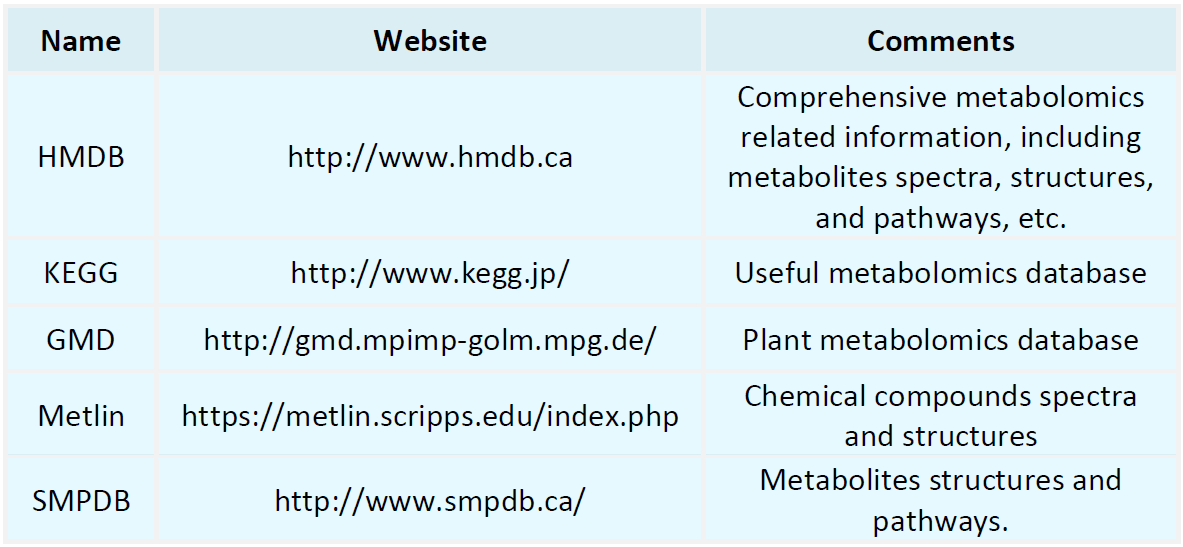
-
• Comprehensive Overview of Mass Spectrometry-Based Protein Phosphorylation Analysis Technologies
Protein phosphorylation is one of the most prevalent and functionally significant post-translational modifications, playing essential roles in a wide range of biological processes, including cell proliferation, differentiation, metabolism, and apoptosis. Precise identification and quantitative characterization of phosphorylation sites enable researchers to systematically decipher the dynamic regulation of cellular signaling networks, elucidate molecular mechanisms underlying disease pathogenesis, and ......
-
• How to Integrate Spatial Proteomics with Multi-Omics Approaches?
The rapid advances in single-cell omics, spatial transcriptomics, and high-throughput mass spectrometry have propelled life-science research into an era of high-dimensional data integration. Conventional proteomics captures changes in protein abundance, yet it often cannot address a fundamental question: where, precisely, are these proteins located within tissues or cells? Spatial proteomics has been developed to fill this gap by enabling visualization and quantitative profiling of protein distributio......
-
• Common Types of Histone Modifications and Their Mass Spectrometric Signatures
Histones are core structural components of chromatin that not only facilitate DNA packaging and maintain chromatin architecture, but also play essential regulatory roles in gene expression, DNA repair, and chromatin remodeling through a wide range of post-translational modifications (PTMs). A broad spectrum of histone modifications has been identified, among which acetylation, methylation, phosphorylation, and ubiquitination are the most extensively studied. Collectively, these modifications constitut......
-
• LC-MS/MS-Based Quantitative Ubiquitinomics
Ubiquitination is a critical post-translational protein modification involved in cellular regulation and participates in numerous essential physiological processes, including protein degradation, signal transduction, cell cycle control, and DNA damage response. However, the accurate and sensitive quantification of ubiquitinated proteins in complex biological systems remains highly challenging. LC-MS/MS (liquid chromatography–tandem mass spectrometry) has become a core analytical platform for the quant......
-
• Role of Protein Ubiquitination in Cellular Signaling Pathways
Within the highly complex cellular regulatory network, protein ubiquitination is not only a fundamental mechanism for maintaining protein homeostasis but also serves as a precise regulatory system in diverse signaling pathways. By covalently attaching one or more ubiquitin molecules to specific substrate proteins, ubiquitination determines the fate of target proteins, whether they undergo activation, inhibition, subcellular relocalization, or complete proteasomal degradation. Particularly in cellular ......
-
• Impact of Ubiquitination on Protein Degradation Pathways
Proteins within cells are not permanent entities after synthesis. Once they have fulfilled their biological functions, they must be promptly recognized and degraded to maintain protein homeostasis and intracellular equilibrium. Cellular proteins are constantly maintained in a dynamic balance between synthesis and degradation, a process that is essential for sustaining proteostasis and normal cellular function. As a highly conserved post-translational modification, ubiquitination precisely labels prote......
-
• Mass Spectrometry-Based Subcellular Proteomics: Advantages, Challenges, and Applications
Proteomics provides a global overview of protein expression at the whole-cell level; however, it lacks the ability to resolve the precise intracellular localization of proteins. In fact, protein function is tightly associated with its subcellular localization. Organelles such as mitochondria, endoplasmic reticulum, Golgi apparatus, and nucleus each perform distinct biological functions, and protein mislocalization often indicates cellular dysfunction and even disease onset. Subcellular proteomics, par......
-
• Why Is Shotgun Proteomics the Preferred Strategy for Comprehensive Proteome Profiling?
In proteomics research, the efficient characterization of protein composition and dynamic changes in complex biological samples remains a fundamental challenge. Owing to its high throughput, broad proteome coverage, and superior analytical sensitivity, shotgun proteomics has emerged as the most widely adopted strategy for comprehensive proteome profiling. It has been extensively applied in studies of disease mechanisms, biomarker discovery, and integrative multi-omics analyses. What Is Shotgun Proteo......
-
• How to Perform Qualitative Shotgun Protein Identification Using DDA Mode?
In life science research, proteins serve as a central link between genomic information and cellular functions. To systematically identify and characterize the full complement of proteins in biological samples, shotgun protein identification has become one of the most widely adopted strategies in proteomics. This approach involves the direct introduction of enzymatically digested peptide mixtures into a mass spectrometry system, coupled with data-dependent acquisition (DDA) for high-throughput data col......
-
• Comprehensive Guide to 4D Label-Free Quantitative Proteomics Workflow
In the post-genomic era, proteomics has emerged as a pivotal approach for elucidating biological processes, uncovering disease mechanisms, and discovering biomarkers. Specifically, label-free quantitative proteomics (LFQ) has gained widespread application in both basic and translational biomedical research due to its high throughput and the advantage of eliminating the need for costly labeling reagents. With the rapid advancement of mass spectrometry technologies, 4D-based label-free quantification pl......
How to order?







