How to Perform Mitochondrial Proteomics by LC-MS/MS and What Workflows Are Used?
-
Differential centrifugation: suitable for routine sample preparation; technically straightforward but typically provides moderate purity.
-
Density gradient centrifugation (e.g., Percoll or sucrose gradients): yields mitochondria with higher purity and is suitable for high-sensitivity downstream mass spectrometry analysis.
-
Commercial isolation kits (e.g., Thermo, Sigma): provide standardized protocols with good reproducibility and are suitable for high-throughput experiments.
-
All buffers should be pre-cooled prior to the experiment, and protease inhibitors should be added.
-
Mitochondrial marker proteins such as Cox IV and VDAC can be used to verify mitochondrial purity (e.g., by Western blotting or LC-MS-based verification prior to proteomic analysis).
-
RIPA or urea-based lysis buffers are suitable for total protein extraction.
-
If membrane proteins are the primary focus, mild detergents such as Triton X-100 or CHAPS may be incorporated.
-
Either total lysis or fractionated lysis (membrane vs. soluble proteins) may be applied depending on the experimental objective.
- The BCA assay is commonly recommended because it is compatible with most lysis buffers and provides relatively high quantitative accuracy.
-
Filter-aided Sample Preparation (FASP): suitable for samples containing strong detergents.
-
In-solution digestion: appropriate when sample quantities are sufficient and buffer conditions are relatively mild.
-
Enzymatic digestion strategies: trypsin single digestion or combined trypsin/LysC digestion can increase protein sequence coverage.
- C18 solid-phase extraction cartridges are commonly used to remove salts and contaminants, thereby improving mass spectrometry signal quality.
-
Orbitrap Fusion Lumos / Exploris series
-
timsTOF Pro 2 (particularly suitable for deep acquisition in DIA mode)
-
DDA (Data-Dependent Acquisition) is commonly used for spectral library generation.
-
DIA (Data-Independent Acquisition) is well suited for high-throughput quantitative analysis and large cohort studies.
-
Nano-liquid chromatography (nanoLC) coupled with long gradient separation improves the resolution of complex peptide mixtures.
-
Dynamic exclusion settings, collision energy, and scan speed should be optimized according to the specific instrument platform.
-
Raw data processing (e.g., MaxQuant, Spectronaut, DIA-NN)
-
Protein identification and quantification
-
Differential analysis (log2 fold change, p-value)
-
Functional enrichment analysis (GO/KEGG, Reactome)
-
Protein-protein interaction (PPI) network analysis (STRING, Cytoscape)
-
Using high-quality mitochondrial protein annotations from the UniProt database can improve annotation accuracy.
-
For membrane proteins or post-translationally modified proteins, dedicated enrichment strategies are often recommended.
-
A dual-platform system based on Orbitrap Exploris 480 and timsTOF Pro 2, supporting both DDA and DIA acquisition modes.
-
Comprehensive mitochondrial isolation services combined with protein purity verification (Western blotting or LC-MS analysis).
-
Support for in-depth characterization of small samples, low-abundance proteins, and post-translationally modified proteins.
-
Integrated bioinformatics reports that facilitate scientific publication and mechanistic investigation.
Mitochondria are central organelles in cellular energy metabolism. In addition to driving ATP production, they also regulate a variety of biological processes, including apoptosis and redox homeostasis. Comprehensive characterization of the mitochondrial proteome is therefore essential for understanding metabolic disorders, cancer, neurodegenerative diseases, and other mitochondria-associated pathologies. Mass spectrometry-based mitochondrial proteomics has become a key strategy for systematically investigating mitochondrial functions.
Overview of the Workflow for Mitochondrial Proteomic Analysis
Mitochondrial proteomic analysis generally involves the following five core steps:
1. Mitochondrial isolation and purification
2. Protein extraction and quantification
3. Protein digestion and peptide purification
4. Mass spectrometry analysis
5. Data analysis and functional annotation
Detailed Experimental Workflow
1. Mitochondrial Isolation and Purification: Ensuring High Purity Is Essential
The quality of mitochondrial isolation directly determines the reliability of downstream proteomic data.
(1) Common Isolation Methods
(2) Optimization Recommendations
2. Protein Extraction and Quantification: Balancing Coverage and Stability
Because mitochondrial membranes possess complex structures and contain diverse protein classes, extraction strategies should balance solubilization efficiency with protein stability.
(1) Extraction Strategies
(2) Protein Quantification
3. Protein Digestion and Peptide Purification: Ensuring Peptide Uniformity and MS Compatibility
(1) Common Methods
(2) Peptide Purification
4. Mass Spectrometry Analysis: High-Resolution Platforms Improve Identification Depth
(1) Common Mass Spectrometry Platforms
(2) DDA vs DIA
(3) Key Parameter Considerations
5. Data Analysis: From Peptides to Biological Interpretation
(1) Analysis Workflow
(2) Important Considerations
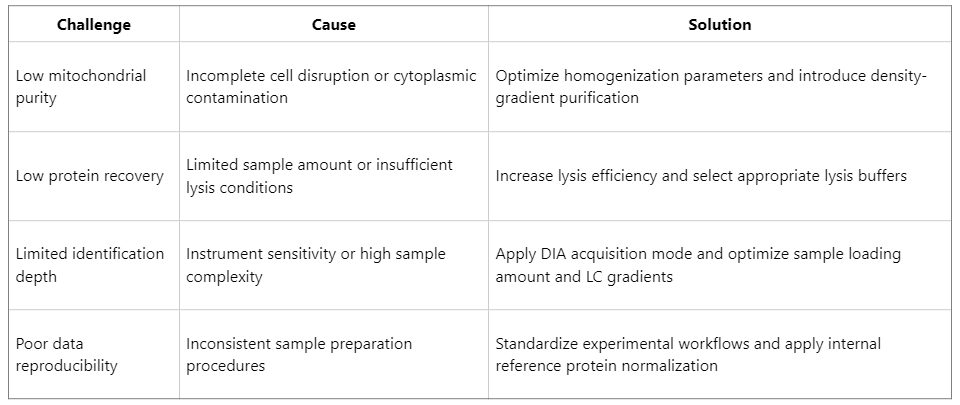
Common Challenges and Optimization Strategies
Advantages of the Mass Spectrometry Platform at MtoZ Biolabs
At MtoZ Biolabs, we have established a highly standardized yet flexible technical workflow specifically designed for mitochondrial proteomic analysis:
Mitochondrial proteomic analysis is a systematic process involving multiple complex steps. Its successful implementation depends on high-quality sample preparation, appropriate selection of mass spectrometry platforms, and rigorous data analysis pipelines. Through well-designed experimental strategies and professional technological support, both protein identification depth and data reliability can be significantly improved, providing robust technical support for research on mitochondrial-related diseases. If you are conducting studies on mitochondrial function, investigating mechanisms of metabolic disorders, or screening potential drug targets, MtoZ Biolabs welcomes your inquiries. Our professional technologies and flexible solutions are dedicated to supporting your research projects.
MtoZ Biolabs, an integrated chromatography and mass spectrometry (MS) services provider.
Related Services
How to order?







