Resources
Proteomics Databases
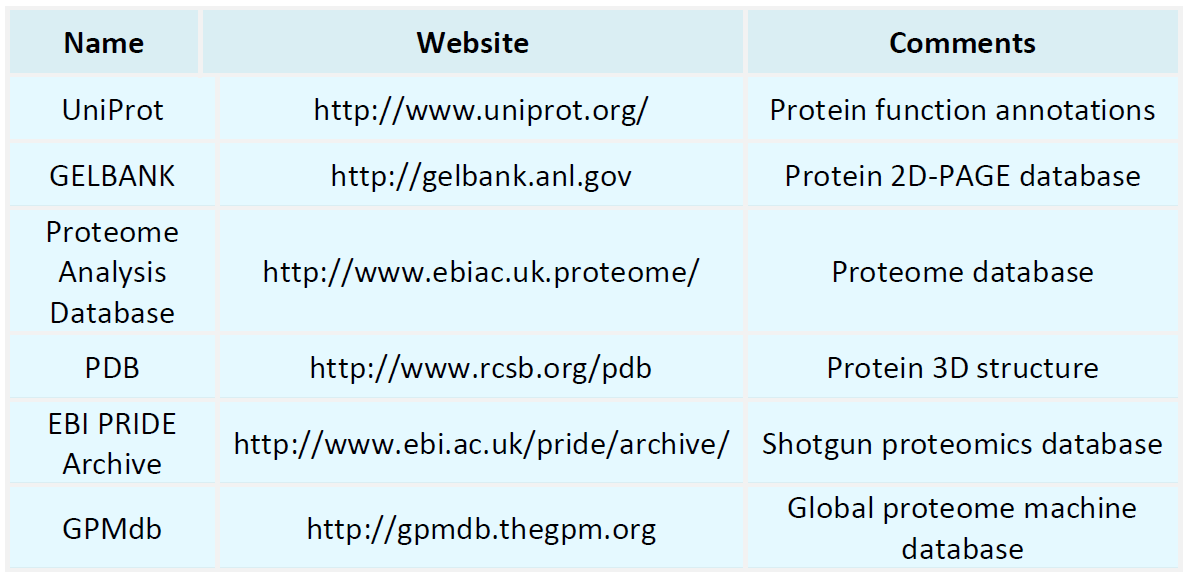
Metabolomics Databases
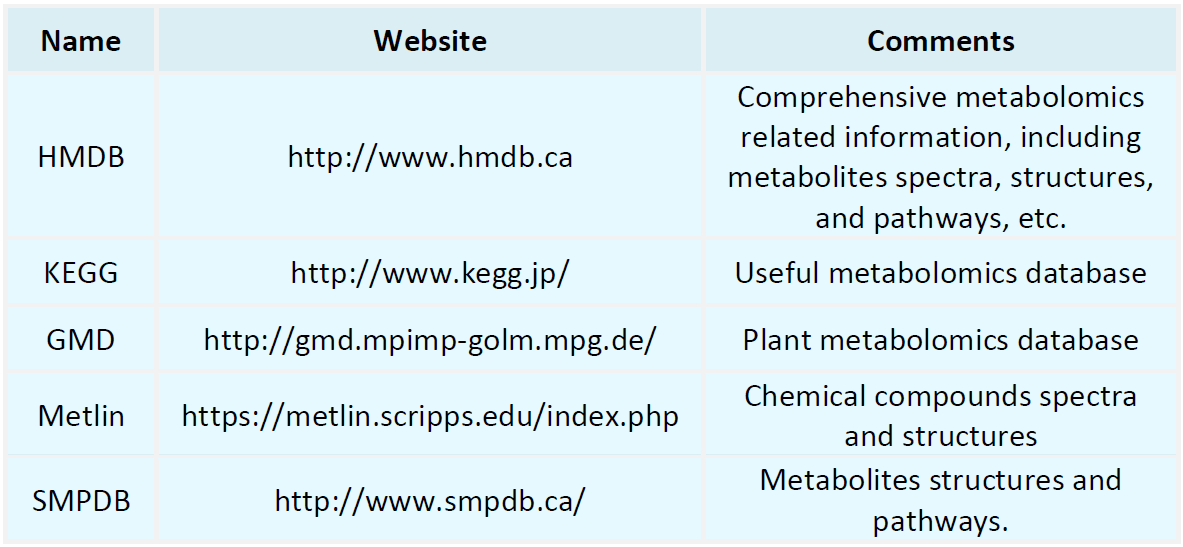
-
Peptide sequencing is a fundamental biochemical method designed to precisely determine the order of amino acids within a peptide chain. Peptides, the essential building blocks of proteins, are short chains of amino acids linked by peptide bonds. These small proteins have diverse biological functions, including roles in hormone signaling and enzymatic catalysis. Accurate peptide sequencing allows researchers to investigate the specific mechanisms, structural characteristics, and biological roles of the......
-
• How is Peptide Sequencing Performed?
Peptide sequencing serves as a critical technique for uncovering the structural and functional information encoded within polypeptides, which are essential to various biological processes. Polypeptides, formed by multiple amino acids linked via peptide bonds, require sequencing to identify their precise amino acid arrangement. This sequencing is analogous to solving a complex puzzle, where each method leverages unique chemical and physical principles to determine amino acid order. For instance, mass......
-
• In What Ways is Mass Spectrometry Employed in Proteomics Research?
Mass spectrometry (MS) is a cornerstone of proteomics research, enabling comprehensive analysis of protein expression, modifications, and interactions within biological systems. Its applications extend beyond individual protein characterization to encompass global insights into proteomic dynamics. Key areas of application include: Proteome Quantification MS provides quantitative insights into protein expression across entire proteomes, offering a comprehensive view of protein abundance in cells, tissu......
-
• What Makes ESI MS/MS an Optimal Approach for Protein Identification?
Electrospray ionization tandem mass spectrometry (ESI MS/MS) is a powerful analytical method that combines electrospray ionization (ESI) and tandem mass spectrometry (MS/MS). It has become a preferred choice for protein identification due to its unique advantages across multiple dimensions: Gentle Ionization for Biomolecules ESI employs a soft ionization process where sample solutions are nebulized into charged droplets under an electric field and auxiliary gas flow. As these droplets evaporate, small......
-
• How is Data Generated by Protein Mass Spectrometry Analyzed and Interpreted?
Protein mass spectrometry (MS) data analysis involves separating, identifying, and quantifying proteins to gain insights into their molecular characteristics. MS measures the mass-to-charge ratio (m/z) and intensity of ionized protein fragments, providing valuable information about molecular weight, sequence, post-translational modifications, and relative abundance. The typical workflow for protein MS data analysis includes the following steps: 1. Sample Preparation Proteins are enzymatically digest......
-
• What is the Typical Amount of Protein Sample Required for Mass Spectrometry Analysis?
Mass spectrometry analysis is a widely applied technique for determining the mass-to-charge ratio (m/z) of ions, particularly in life sciences. By ionizing sample components and separating the resulting ions based on their m/z, it generates spectra that reveal molecular mass, protein sequences, and post-translational modifications. Its versatility and precision make mass spectrometry analysis indispensable in proteomics and related fields. The protein sample required for mass spectrometry analysis v......
-
• By What Mechanisms Does Mass Spectrometry Facilitate Protein Identification?
Mass spectrometry (MS) is a powerful analytical tool for identifying and characterizing proteins. It integrates sample preparation, peptide analysis, and data interpretation to provide comprehensive insights into protein structure and function. Protein Sample Preparation Proteins are extracted from biological sources, such as cells, tissues, or fluids, using methods that preserve their integrity. For instance, cell lysates containing detergents and protease inhibitors efficiently release proteins. T......
-
• What is Quantitative Phosphoproteomics?
Quantitative phosphoproteomics is a branch of bioinformatics dedicated to studying protein phosphorylation levels and their role in regulating dynamic biological processes such as cell signaling, gene expression, and cell cycle progression. Advances in proteomics and computational techniques have expanded its applications across various areas of biological research. Protein phosphorylation is a common post-translational modification in which a phosphate group is added to specific amino acid residues......
-
• What Does Protein Sequencing by Mass Spectrometry Entail?
Protein sequencing by mass spectrometry is a cutting-edge analytical method used to determine the amino acid sequences of proteins with high precision. In modern biological research, proteins serve as the primary executors of cellular functions, making their structural and functional studies a cornerstone of scientific inquiry. Proteins participate in nearly all cellular biochemical reactions and play key roles in intercellular signaling, material transport, and immune defense. Understanding protein s......
-
• Can You Provide Representative Examples of Peptide Sequences?
Peptide sequences are central to numerous analytical applications in biotechnology and life sciences. Peptide mapping, a powerful technique, allows detailed analysis of protein structure, sequence integrity, and modifications. Below are representative applications highlighting the versatility of peptide sequences in research and development. 1. Quality Control of Recombinant Protein Drugs In the production of recombinant human interferon, peptide mapping ensures the accuracy and completeness of the am......
How to order?







